Microbial Metabolism 1
Bạn đang xem bản rút gọn của tài liệu. Xem và tải ngay bản đầy đủ của tài liệu tại đây (1 MB, 45 trang )
Microbial Metabolism
Microbiology 274
March 22, 2007
Free energy of reactions
Temperature
∆G = ∆H - T∆S
Free Energy:
How much energy
during a reaction is
available to do work
Entropy:
Enthalpy:
How much heat is
lost or gained during
the reaction
How much
randomness is lost
or gained during the
reaction
Page 152 in your text
Free energy of reactions
Exergonic reaction:
∆G°’ is negative
A+B
C+D
Endergonic reaction: ∆G°’ is positive
A+B
C+D
Figure 8.5
Metabolism
Organisms need to synthesize many complex organic molecules
Synthesis from monomers to polymers -- these reactions are
endergonic and require a source of energy
ATP
ATP is used as a source of energy to
change endergonic reactions to
exergonic reactions
Energy Coupling
A+B
C+D
ATP
A+B
ADP + Pi
C +enzyme
D
Figure 8.6
Enzymes as catalysts for
reactions
Enzymes are proteins
Highly specific for the reaction that they catalyze
Reduce the energy of activation (E ) of a reaction
a
C6H12O6 + O2
CO2 + H2O
Ea
∆G°’
Figure 8.14 is similar
Back to ATP…
ATP is constantly depleted by cells
ADP + P = ATP
i
∆G = 7.3 kcal/mole
In order to make more ATP, cells need to input energy…
Oxidation-Reduction Reactions
Loss of Electrons = Oxidation (LEO)
Gain of Electrons = Reduction (GER)
Electron Donors and Electron Acceptors
Acceptor + ne-
Donor
Page 153 in your text
Standard Reduction Potential
Measure of the tendency of a donor to lose electrons (E’ )
0
2H+ + 2eNAD+ + 2H+ + 2e-
H2
E’0 = -420 mV
NADH + H2 E’0 =
-320 mV
1
/2O2 + 2H+ + 2emV
H 2O
E’0 = +820
Figure 8-7
Mechanisms of Energy Release
Fermentation -- oxidation of an organic compound in the absence of
external electron acceptor
Respiration -- oxidation of an organic compound where oxygen is the
final electron acceptor
Anaerobic Respiration -- oxidation of organic compounds where an
external substrate other than oxygen serves as final electron
acceptor
The Three Ways to Make ATP
Substrate Level Phosphorylation
Glycolysis
Fermentation
Respiration
Electron
transport systems
Photophosphorylation
Photosynthesis
Chapter 9 in your text
Substrate-Level Phosphorylation
Glycolysis
Glucose + 2ADP + 2Pi + 2NAD+
2 Pyruvate + 2ATP + 2NADH + 2H+
Glycolysis (continued)
Glucose
ATP
ADP
6 Carbon
Stage
hexokinase
Glucose 6-phosphate
phosphoglucoisomerase
Fructose 6-phosphate
ATP
Fructose 1,6-bisphosphate
phosphofructokinase
ADP
aldolase
Glyceraldehyde 3- P
Dihydroxyacetone- P
Glycolysis (continued)
Glyceraldehyde 3- P
Pi
1,3-Bisphosphoglycerate
Dihydroxyacetone- P
NAD+
NADH
+ H+
ADP
3-Phosphoclycerate
2-Phosphoglycerate
dehydrogenase
ATP
phosphoglycerate
kinase
phosphoglycerate
mutase
3 Carbon
Stage
Glycolysis (continued)
2-Phosphoglycerate
enolase
Phosphoenolpyuvate
ADP
ATP
Pyruvate
pyruvate
kinase
3 Carbon
Stage
Conversion of glucose to pyruvate results in the formation of NADH
To maintain homeostasis, cells must reoxidize NADH to NAD+ or glycolysis
will stop!
Cells use electron acceptors to oxidize NADH
Fermentation -- reduction of organic compounds
Electron transport chains -- reduction of O2
Fermentation
NADH
NAD+
pyruvate
lactate
lactate
dehydrogenase
e.g. Bacillus
Lactobacillus
Streptococcus
Fermentation (continued)
CO2
pyruvate
pyruvate
decarboxylase
NADH
acetaldehyde
NAD+
ethanol
alcohol
dehydrogenase
Eg. Yeast
Diversity of microbial fermentation
Pathway
End Products
Examples
Lactic Acid
Lactic acid (2 molecules)
Lactobacillus, Enterococcus,
Streptococcus spp.
Heterolactic
Lactic acid, ethanol and CO2
Leuconostoc
Alcohol
Ethanol and CO2
Saccharomyces (yeast)
Proprionic acid
Proprionic acid and CO2
Proprionibacterium spp.
Butyric acid
Butyric acid, butanol, acetone,
isopropyl alcohol and CO2
Clostridium spp.
Butanediol
Butanediol and CO2
Enterobacter, Serratia, Erwinia,
and Klebsiella
Mixed acid
Ethanol, acetic acid, lactic acid,
E. coli, Salmonella, and Shigella
succinic acid, formic acid and CO2
Methanogenesis Methane and CO2
Archaea
Respiration
NADH can be reoxidized by donating electrons to an external electron acceptor
such as oxygen.
The lower the redox potential of the acceptor, the more energy can be obtained
in the form of ATP.
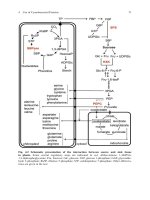
Tricarboxylic Acid Cycle
Lipid and protein
metabolites
malate
dehydrogenase
pyruvate
NAD+
acetyl CoA
NADH + H+
oxaloacetatecitrate
citrate
synthase
L-malate
isocitrate
NADH
fumarate
isocitrate
dehydrogenase
FADH2
αketoglutarat
e α-ketoglutarate
succinate
succinate
dehydrogenase
succinyl CoA
dehydrogenase
Tricarboxylic Acid Cycle
pyruvate
- acetyl CoA
-
NAD+
ATP, acetyl CoA, NADH
malate
dehydrogenase
oxaloacetatecitrate
NADH + H+
ATP, succinyl CoA, citrate, NAD+
citrate
synthase
L-malate
isocitrate
NADH
fumarate
-
ATP
isocitrate
dehydrogenase
FADH2
αsuccinyl CoA
ketoglutarat
NADH
e α-ketoglutarate
-
succinate
succinate
dehydrogenase
succinyl CoA
dehydrogenase
Tricarboxylic Acid Cycle
AMP, CoA, NAD+
pyruvate
NAD+
acetyl CoA
NADH + H+
+
+ ADP
malate
dehydrogenase
oxaloacetatecitrate
citrate
synthase
L-malate
isocitrate
NADH
fumarate
+ ADP
isocitrate
dehydrogenase
FADH2
αketoglutarat
e α-ketoglutarate
succinate
succinate
dehydrogenase
succinyl CoA
dehydrogenase